Fuzzy complex
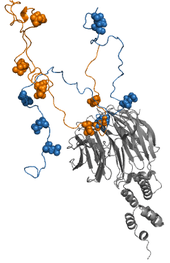
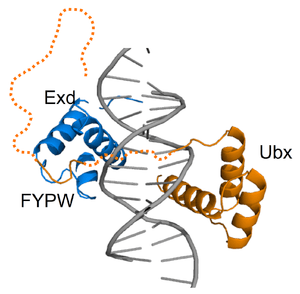
Fuzzy complexes are protein complexes, where structural ambiguity or multiplicity exists and is required for biological function.[1][2] Alteration, truncation or removal of conformationally ambiguous regions impacts the activity of the corresponding complex.[3][4][5] Fuzzy complexes are generally formed by intrinsically disordered proteins.[6][7] Structural multiplicity usually underlies functional multiplicity of protein complexes [8][9][10] following a fuzzy logic. Distinct binding modes of the nucleosome are also regarded as a special case of fuzziness.[11][12]
Historical background
For almost 50 years molecular biology was based on two dogmas: (i) equating biological function of the protein with a unique three-dimensional structure and (ii) assuming exquisite specificity in protein complexes. Specificity/selectivity is ensured by unambiguous set of interactions formed between the protein and its ligand (another protein, DNA, RNA or small molecule). Many protein complexes however, contain functionally important/critical regions, which remain highly dynamic in the complex or adopt different conformations.[13] This phenomenon is defined fuzziness. The most pertinent example is the cyclin-dependent kinase inhibitor Sic1, which binds to the SCF subunit of Cdc4 in a phosphorylation dependent manner.[14] No regular secondary structures are gained upon phosphorylation and the different phosphorylation sites interchange in the complex.[15]
Classification of fuzzy complexes
Structural ambiguity in protein complexes covers a wide spectrum.[1] In a polymorphic complex, the protein adopts two or more different conformations upon binding to the same partner, and these conformations can be resolved.[16] Clamp,[17] flanking [18][19] and random complexes [20][21] are dynamic, where ambiguous conformations interchange with each other and cannot be resolved. Interactions in fuzzy complexes are usually mediated by short motifs,.[22][23] Flanking regions are tolerant to sequence changes as long as the amino acid composition is maintained, for example in case of linker histone C-terminal domains [24] and H4 histone N-terminal domains.[25]
Regulatory pathways via fuzzy regions
Fuzzy regions modulate the conformational equilibrium [26] or flexibility [3][27] of the binding interface via transient interactions.[28] Dynamic regions can also compete with binding sites [29] or tether them to the target.[30] Modifications of fuzzy regions by further interactions,[8][31] or posttranslational modifications [32][33] impact binding affinity or specificity. Alternative splicing can modulate the length of fuzzy regions resulting in context-dependent binding (e.g. tissue-specificity) on the complex.[34][35][36] EGF/MAPK, TGF-β and WNT/Wingless signaling pathways employ tissue-specific fuzzy regions.
References
- 1 2 Tompa, P. & Fuxreiter, M. (Jan 2008) "Fuzzy complexes: polymorphism and structural disorder in protein-protein interactions". Trends Biochem Sci 33,(1): 2-8. PMID 18054235.
- ↑ Fuxreiter, M. & Tompa, P. (2011) Fuzziness: Structural Disorder in Protein Complexes Austin, New York.
- 1 2 Pufall, M.A., Lee, G.M., Nelson, M.L., Kang, H.S., Velyvis, A. et al. (Jul 1 2005) "Variable control of Ets-1 DNA binding by multiple phosphates in an unstructured region". Science 309,(5731): 142-5. PMID 15994560.
- ↑ Bhattacharyya, R.P., Remenyi, A., Good, M.C., Bashor, C.J., Falick, A.M. et al. (Feb 10 2006) "The Ste5 scaffold allosterically modulates signaling output of the yeast mating pathway". Science 311,(5762): 822-6. PMID 16424299.
- ↑ Liu, Y., Matthews, K.S. & Bondos, S.E. (Jul 24 2009) "Internal regulatory interactions determine DNA binding specificity by a Hox transcription factor". J Mol Biol 390,(4): 760-74. doi: S0022-2836(09)00629-9 [pii]
- ↑ Romero, P., Obradovic, Z., Kissinger, C.R., Villafranca, J.E., Garner, E. et al. 1998) "Thousands of proteins likely to have long disordered regions". Pac. Symp. Biocomputing. 3: 437-448. PMID 9697202.
- ↑ Wright, P.E. & Dyson, H.J. 1999) "Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm". J Mol Biol 293,(2): 321-31. PMID. 10550212
- 1 2 Galea, C.A., Nourse, A., Wang, Y., Sivakolundu, S.G., Heller, W.T. et al. (Feb 22 2008) "Role of intrinsic flexibility in signal transduction mediated by the cell cycle regulator, p27 Kip1". J Mol Biol 376,(3): 827-38. PMID 18177895.
- ↑ Fuxreiter, M., Tompa, P., Simon, I., Uversky, V.N., Hansen, J.C. et al. (Dec 2008) "Malleable machines take shape in eukaryotic transcriptional regulation". Nat Chem Biol 4,(12): 728-37. doi: nchembio.127 [pii] 10.1038/nchembio.127. PMID 19008886.
- ↑ Wang, Y., Fisher, J.C., Mathew, R., Ou, L., Otieno, S. et al. (April 2011) "Intrinsic disorder mediates the diverse regulatory functions of the Cdk inhibitor p21". Nat. Chem. Biol. 7: 214-221. PMID 21358637.
- ↑ Belch, Y., Yang, J., Liu, Y., Malkaram, S.A., Liu, R. et al. 2010) "Weakly positioned nucleosomes enhance the transcriptional competency of chromatin". PLoS ONE 5,(9): e12984. doi: 10.1371/journal.pone.0012984. PMID 20886052.
- ↑ Tsui, K., Dubuis, S., Gebbia, M., Morse, R.H., Barkai, N. et al. (Nov 2011) "Evolution of nucleosome occupancy: conservation of global properties and divergence of gene-specific patterns". Mol Cell Biol 31,(21): 4348-55. doi: MCB.05276-11 [pii] 10.1128/MCB.05276-11. PMID 21896781.
- ↑ Fuxreiter, M. (Jan 2012) "Fuzziness: linking regulation to protein dynamics". Mol Biosyst 8,(1): 168-77. doi: 10.1039/c1mb05234a. PMID 21927770.
- ↑ Nash, P., Tang, X., Orlicky, S., Chen, Q., Gertler, F.B. et al. (Nov 29 2001) "Multisite phosphorylation of a CDK inhibitor sets a threshold for the onset of DNA replication". Nature 414,(6863): 514-21. doi. PMID 11734846.
- ↑ Mittag, T., Orlicky, S., Choy, W.Y., Tang, X., Lin, H. et al. (Nov 18 2008) "Dynamic equilibrium engagement of a polyvalent ligand with a single-site receptor". Proc Natl Acad Sci U S A 105,(46): 17772-7. doi: 0809222105 [pii] 10.1073/pnas.0809222105. PMID 19008353.
- ↑ Didry, D., Cantrelle, F.X., Husson, C., Roblin, P., Moorthy, A.M. et al. (Dec 23 2011) "How a single residue in individual beta-thymosin/WH2 domains controls their functions in actin assembly". EMBO J. doi: emboj2011461 [pii] 10.1038/emboj.2011.461. PMID 22193718.
- ↑ Fontes, M.R., Teh, T. & Kobe, B. (Apr 14 2000) "Structural basis of recognition of monopartite and bipartite nuclear localization sequences by mammalian importin-alpha". J Mol Biol 297,(5): 1183-94. PMID 10764582.
- ↑ Zor, T., Mayr, B.M., Dyson, H.J., Montminy, M.R. & Wright, P.E. (Nov 1 2002) "Roles of phosphorylation and helix propensity in the binding of the KIX domain of CREB-binding protein by constitutive (c-Myb) and inducible (CREB) activators". J Biol Chem 277,(44): 42241-8. PMID 12196545.
- ↑ Selenko, P., Gregorovic, G., Sprangers, R., Stier, G., Rhani, Z. et al. 2003) "Structural basis for the molecular recognition between human splicing factors U2AF65 and SF1/mBBP". Mol. Cell 11: 965-976. PMID 12718882.
- ↑ Pometun, M.S., Chekmenev, E.Y. & Wittebort, R.J. (Feb 27 2004) "Quantitative observation of backbone disorder in native elastin". J Biol Chem 279,(9): 7982-7. PMID 14625282.
- ↑ Sigalov, A., Aivazian, D. & Stern, L. (Feb 24 2004) "Homooligomerization of the cytoplasmic domain of the T cell receptor zeta chain and of other proteins containing the immunoreceptor tyrosine-based activation motif". Biochemistry 43,(7): 2049-61. PMID 14967045.
- ↑ Neduva, V. & Russell, R.B. (Jun 13 2005) "Linear motifs: evolutionary interaction switches". FEBS Lett 579,(15): 3342-5. PMID 15943979.
- ↑ Davey, N.E., Trave, G. & Gibson, T.J. (Mar 2011) "How viruses hijack cell regulation". Trends Biochem Sci 36,(3): 159-69. doi: S0968-0004(10)00200-8 [pii] 10.1016/j.tibs.2010.10.002. PMID 21146412.
- ↑ Lu, X., Hamkalo, B., Parseghian, M.H. & Hansen, J.C. (Jan 13 2009) "Chromatin condensing functions of the linker histone C-terminal domain are mediated by specific amino acid composition and intrinsic protein disorder". Biochemistry 48,(1): 164-72. doi: 10.1021/bi801636y 10.1021/bi801636y [pii]. PMID 19072710.
- ↑ McBryant, S.J., Klonoski, J., Sorensen, T.C., Norskog, S.S., Williams, S. et al. (Jun 19 2009) "Determinants of histone H4 N-terminal domain function during nucleosomal array oligomerization: roles of amino acid sequence, domain length, and charge density". J Biol Chem 284,(25): 16716-22. doi: M109.011288 [pii] 10.1074/jbc.M109.011288. PMID 19395382.
- ↑ Naud, J.F., McDuff, F.O., Sauve, S., Montagne, M., Webb, B.A. et al. (Sep 27 2005) "Structural and thermodynamical characterization of the complete p21 gene product of Max". Biochemistry 44,(38): 12746-58. doi: 10.1021/bi0500729. PMID 16171389.
- ↑ Lee, G.M., Pufall, M.A., Meeker, C.A., Kang, H.S., Graves, B.J. et al. (Oct 17 2008) "The affinity of Ets-1 for DNA is modulated by phosphorylation through transient interactions of an unstructured region". J Mol Biol 382,(4): 1014-30. doi. PMID 18692067.
- ↑ Fuxreiter, M., Simon, I. & Bondos, S. (Aug 2011) "Dynamic protein-DNA recognition: beyond what can be seen". Trends Biochem Sci 36,(8): 415-23. doi: S0968-0004(11)00059-4 [pii]10.1016/j.tibs.2011.04.006. PMID 21620710.
- ↑ Watson, M., Stott, K. & Thomas, J.O. (Dec 14 2007) "Mapping intramolecular interactions between domains in HMGB1 using a tail-truncation approach". J Mol Biol 374,(5): 1286-97. doi: S0022-2836(07)01287-9 [pii] 10.1016/j.jmb.2007.09.075. PMID 17988686.
- ↑ Olson, K.E., Narayanaswami, P., Vise, P.D., Lowry, D.F., Wold, M.S. et al. (Oct 2005) "Secondary structure and dynamics of an intrinsically unstructured linker domain". J Biomol Struct Dyn 23,(2): 113-24. doi: d=3021&c=4185&p=13080&do=detail [pii]. PMID 16060685.
- ↑ Ahmed, M.A., Bamm, V.V., Shi, L., Steiner-Mosonyi, M., Dawson, J.F. et al. (Jan 2009) "Induced secondary structure and polymorphism in an intrinsically disordered structural linker of the CNS: solid-state NMR and FTIR spectroscopy of myelin basic protein bound to actin". Biophys J 96,(1): 180-91. doi: S0006-3495(08)00040-4 [pii]10.1016/j.bpj.2008.10.003. PMID 19134474.
- ↑ Jonker, H.R., Wechselberger, R.W., Pinkse, M., Kaptein, R. & Folkers, G.E. (Apr 2006) "Gradual phosphorylation regulates PC4 coactivator function". FEBS J 273,(7): 1430-44. doi: EJB5165 [pii] 10.1111/j.1742-4658.2006.05165.x. PMID 16689930.
- ↑ Tsunaka, Y., Toga, J., Yamaguchi, H., Tate, S., Hirose, S. et al. (Sep 4 2009) "Phosphorylated intrinsically disordered region of FACT masks its nucleosomal DNA binding elements". J Biol Chem 284,(36): 24610-21. doi: M109.001958 [pii] 10.1074/jbc.M109.001958. PMID 19605348.
- ↑ Tanaka, T., Kawashima, H., Yeh, E.T. & Kamitani, T. (Aug 29 2003) "Regulation of the NEDD8 conjugation system by a splicing variant, NUB1L". J Biol Chem 278,(35): 32905-13. doi: 10.1074/jbc.M212057200 M212057200 [pii]. PMID 12816948.
- ↑ Liu, Y., Matthews, K.S. & Bondos, S.E. (Jul 25 2008) "Multiple intrinsically disordered sequences alter DNA binding by the homeodomain of the Drosophila hox protein ultrabithorax". J Biol Chem 283,(30): 20874-87. PMID 18508761.
- ↑ Brayer, K.J., Lynch, V.J. & Wagner, G.P. (Aug 9 2011) "Evolution of a derived protein-protein interaction between HoxA11 and Foxo1a in mammals caused by changes in intramolecular regulation". Proc Natl Acad Sci U S A 108,(32): E414-20. doi: 1100990108 [pii]10.1073/pnas.1100990108. PMID 21788518.